[口头报告]基于深度矩阵分解的空间表观基因组数据去噪
基于深度矩阵分解的空间表观基因组数据去噪
编号:70
稿件编号:48 访问权限:仅限参会人
更新:2026-03-23 17:20:37
浏览:50次
口头报告
报告开始:2026年03月27日 15:10 (Asia/Shanghai)
报告时间:20min
所在会议:[S2] “一作面对面”论坛(信息) » [s2] “一作面对面”论坛(信息)
暂无文件
摘要
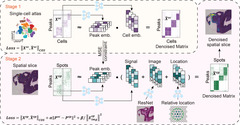
Spatial epigenomics (SE) technologies profile epigenomic landscapes within intact tissues, preserving spatial context and enabling the study of gene regulatory mechanisms in situ. However, current SE datasets typically suffer from low signal detection, substantial noise and extremely sparse peak matrices, which pose considerable challenges for downstream analysis. Here we introduce SPEED (spatial epigenomic data denoising), a deep matrix factorization framework that leverages atlas-level single-cell epigenomic data and spatial context to impute and denoise SE data. In comprehensive benchmarks on both simulated data and real SE tissue datasets, SPEED outperformed five state-of-the-art methods across diverse tissues and technologies. Moreover, SPEED’s denoised outputs facilitated downstream analyses such as differential chromatin accessibility analysis, epigenomic spatial domain identification and gene activity inference. Collectively, our results indicate that SPEED is a generalizable tool for improving data quality and biological insights in SE.
关键字
Spatial epigenomics;Deep matrix factorization;Denoising;Epigenomic spatial domain
报告人

发表评论